Co-translational protein folding in the light of ribosome evolution [EVOCOTF]
In the cell, proteins are synthesised during translation in machines called ribosomes, which use the genomic information coded in mRNA as instructions and, based on them, add one amino acid at a time to the extending nascent polypeptide chain. During this biosynthesis, the nascent chain travels along a long ribosome exit tunnel, and when it reaches its end, it can start to fold into its native shape co-translationally. This process is fundamental for the cells as any problem can drive the protein to misfold, which can cause many disastrous diseases like cystic fibrosis, alpha-1 antitrypsin deficiency or multiple neurodegenerative disorders (Parkinson’s and Alzheimer’s). Despite its importance, much about protein co-translational folding (coTF) is still unknown. The main reason is that it is very difficult to study it experimentally. One of the consequences of these challenges was the development of various computational techniques that can assist experiments in unravelling co-translational protein folding. Molecular dynamics simulations are particularly helpful and have already provided insights into the folding of proteins in E.coli.
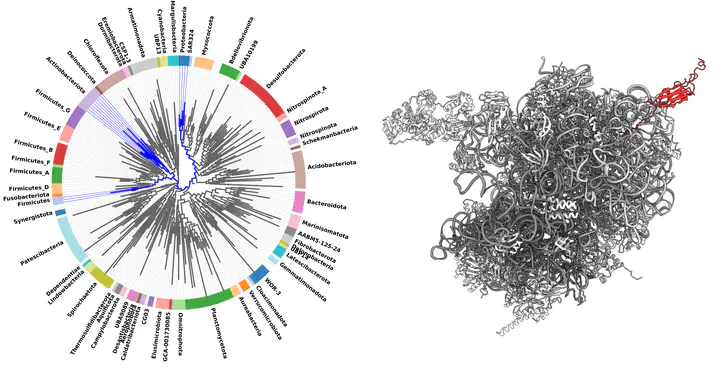
In our project, we will combine molecular dynamics simulations with bioinformatics to extend our understanding of co-translational protein folding, especially in the light of ribosome evolution. We will simulate coTF on various available ribosome structures and analyse how different ribosomes can affect it. Moreover, as ribosome structures have been obtained for only a small fraction of organisms (~40), we will conduct genome-wide bioinformatics analysis to uncover and characterise all proteins that build ribosome exit tunnel in different organisms. Equipped with this extensive data, we will build structural models of some of the most diverse ribosome exit tunnels, which have not yet been studied experimentally. Subsequently, with molecular dynamics simulations, we will study how they impact co-translational protein folding. Finally, we will extend our bioinformatics analysis and uncover all ribosome-associated chaperones in bacteria, a special class of proteins that bind to the ribosome and help proteins to adopt their native shape. By characterising them, we will gain a crucial understanding of the cooperation between the ribosome and chaperones in co-translational protein folding.
Overall, our study will provide insights into one of the most fundamental biological processes impacting every aspect of cell function.
This research is part of the project No. 2022/47/P/NZ1/03127 within the POLONEZ BIS programme co-funded by the National Science Centre and the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement No. 945339


